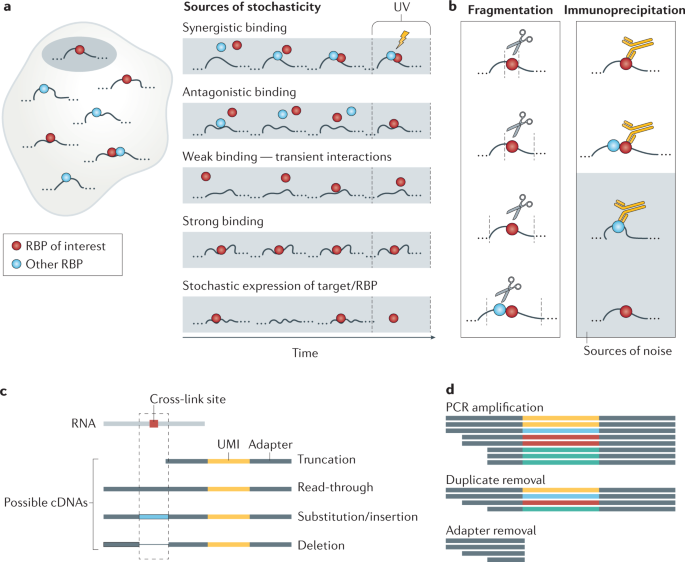
Pervasive and dynamic protein binding sites of the mRNA transcriptome in Saccharomyces cerevisiae | Genome Biology | Full Text

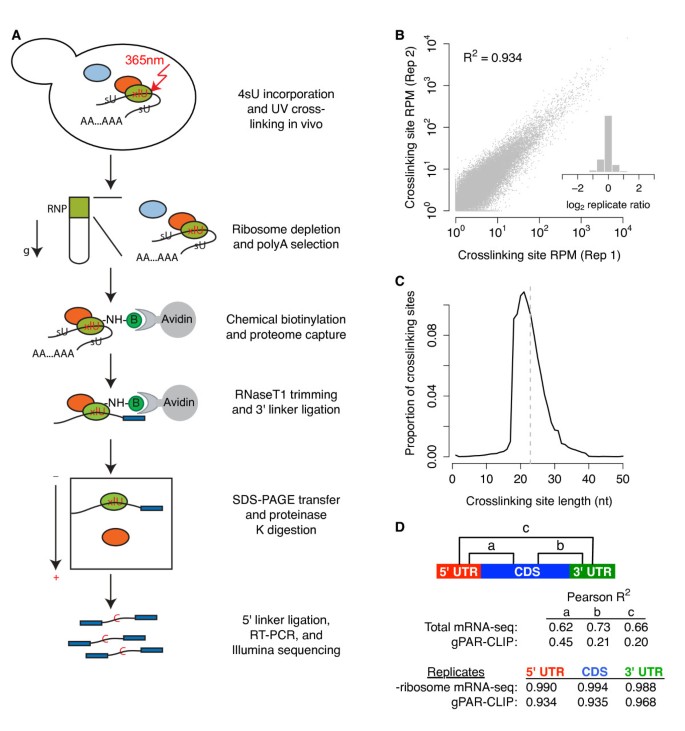
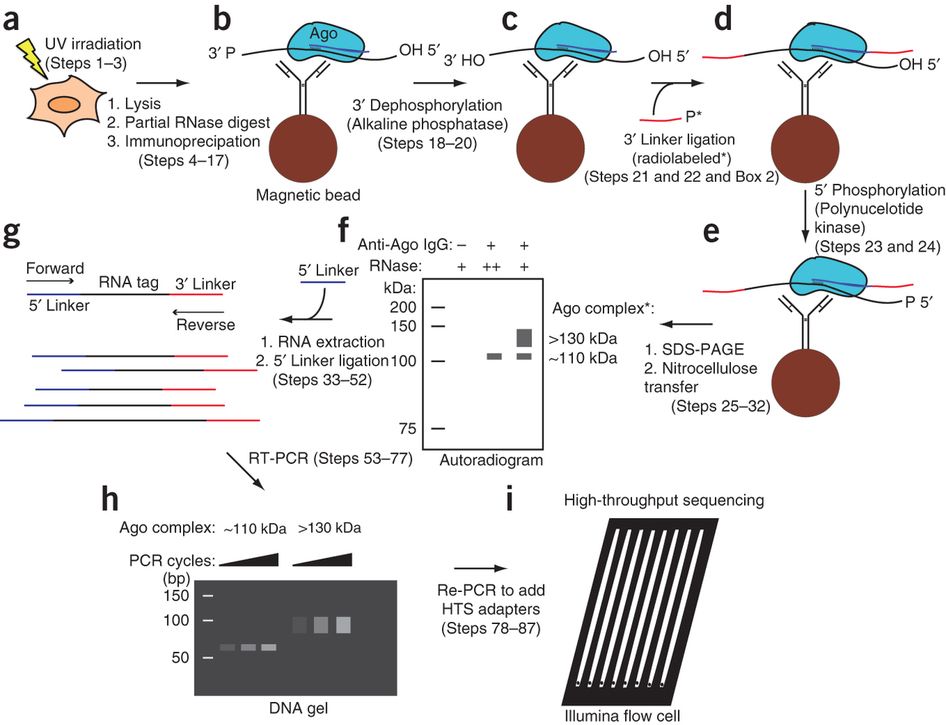
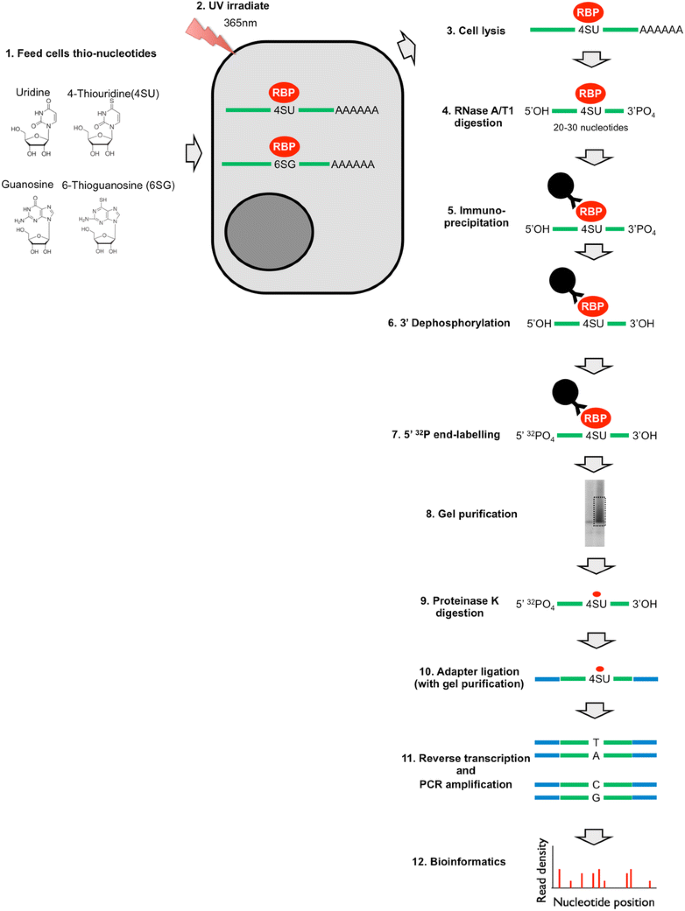
Puf5p HITS-CLIP. (a) Summary of HITS-CLIP protocol. S. cerevisiae cells... | Download Scientific Diagram

fSHAPE, fSHAPE-eCLIP, and SHAPE-eCLIP probe transcript regions that interact with specific proteins - ScienceDirect

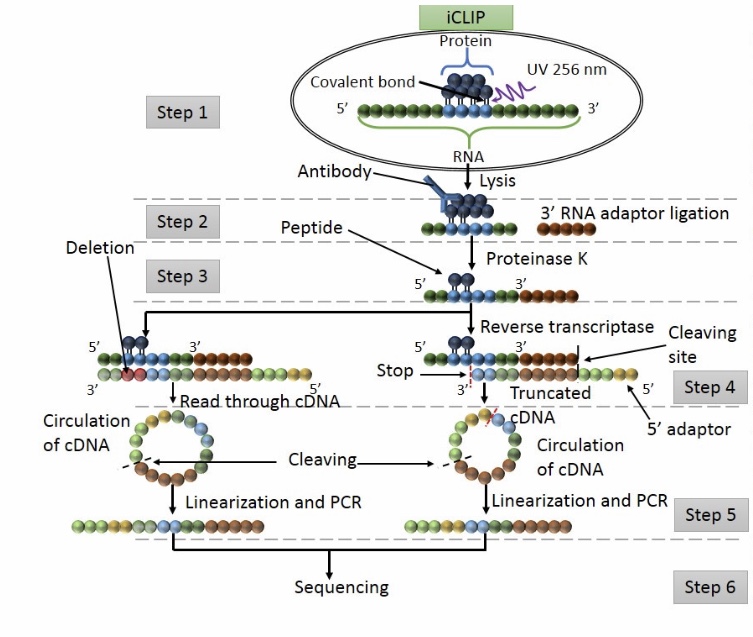
iCLIP - Transcriptome-wide Mapping of Protein-RNA Interactions with Individual Nucleotide Resolution | Protocol

Theoretical and Experimental Background - Bioinformatics Team (BioITeam) at the University of Texas - UT Austin Wikis

Pipeline for extracting MIWI CLIP-Seq features (see Section 2). The RNA... | Download Scientific Diagram

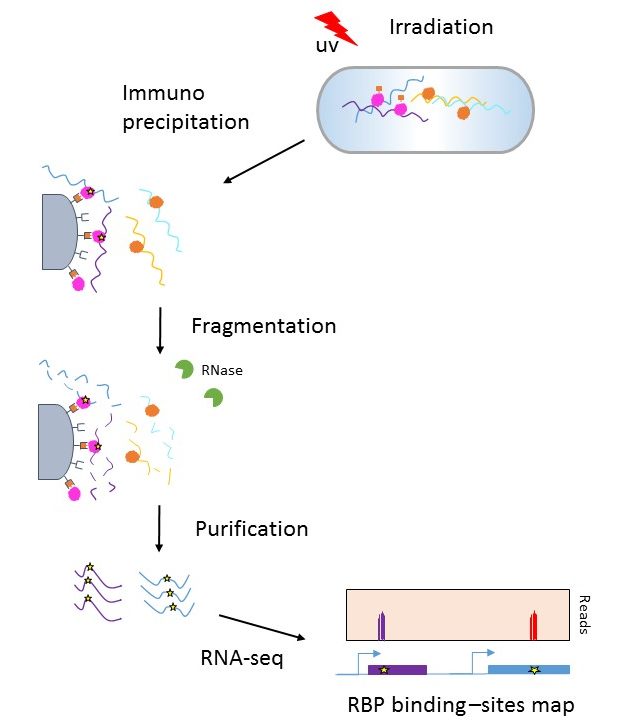
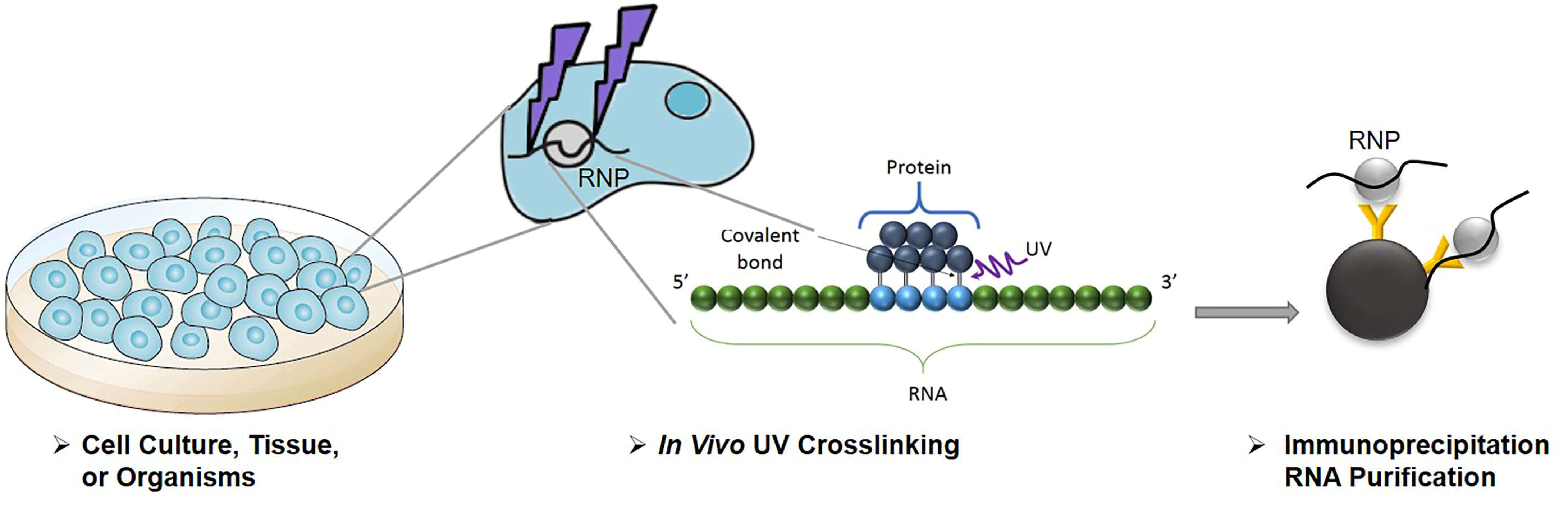
Schematic workflow of optimized CLIP. Cells are irradiated with 254 nm... | Download Scientific Diagram
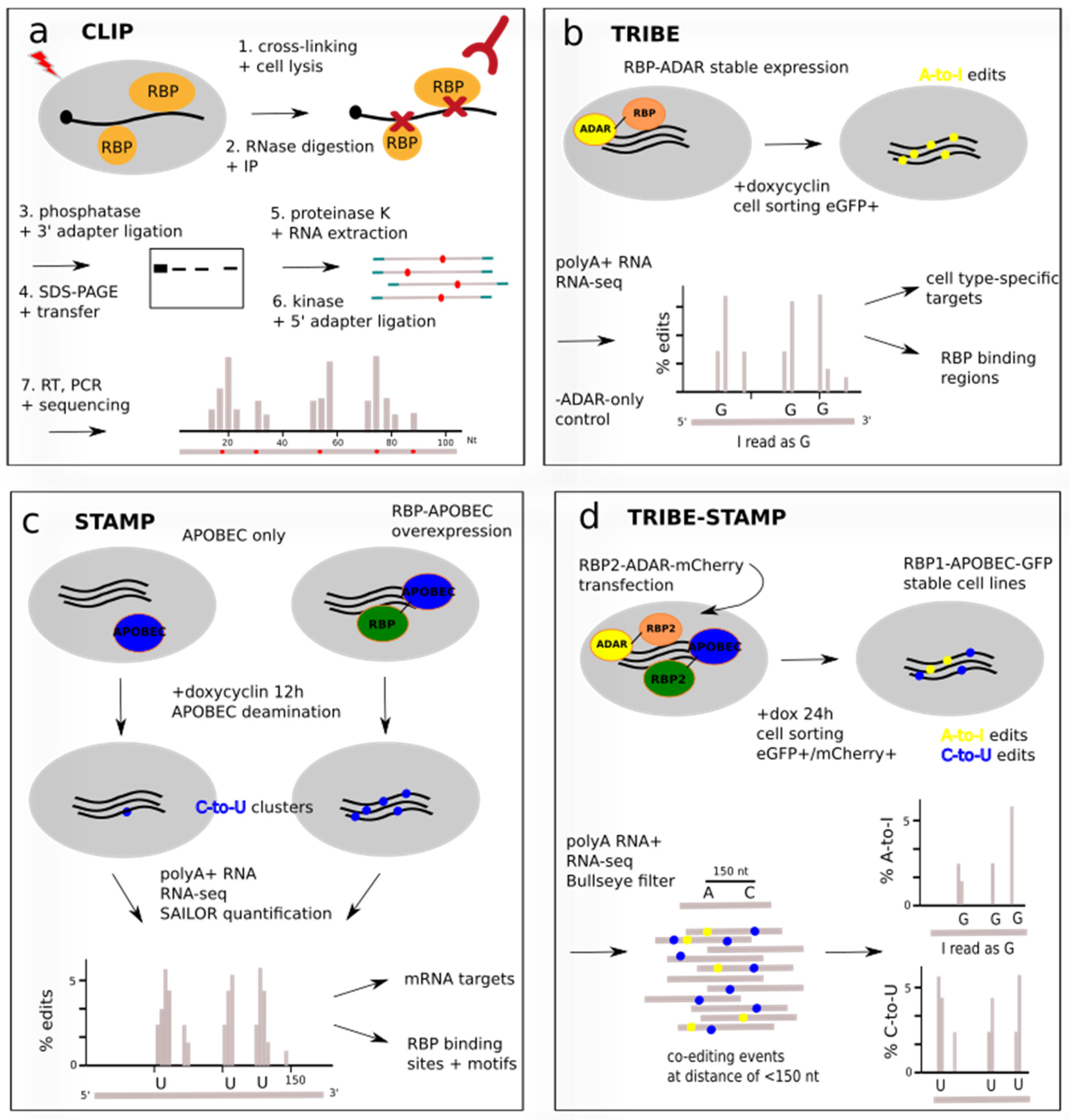
BioChem | Free Full-Text | Current Technical Approaches to Study RNA–Protein Interactions in mRNAs and Long Non-Coding RNAs
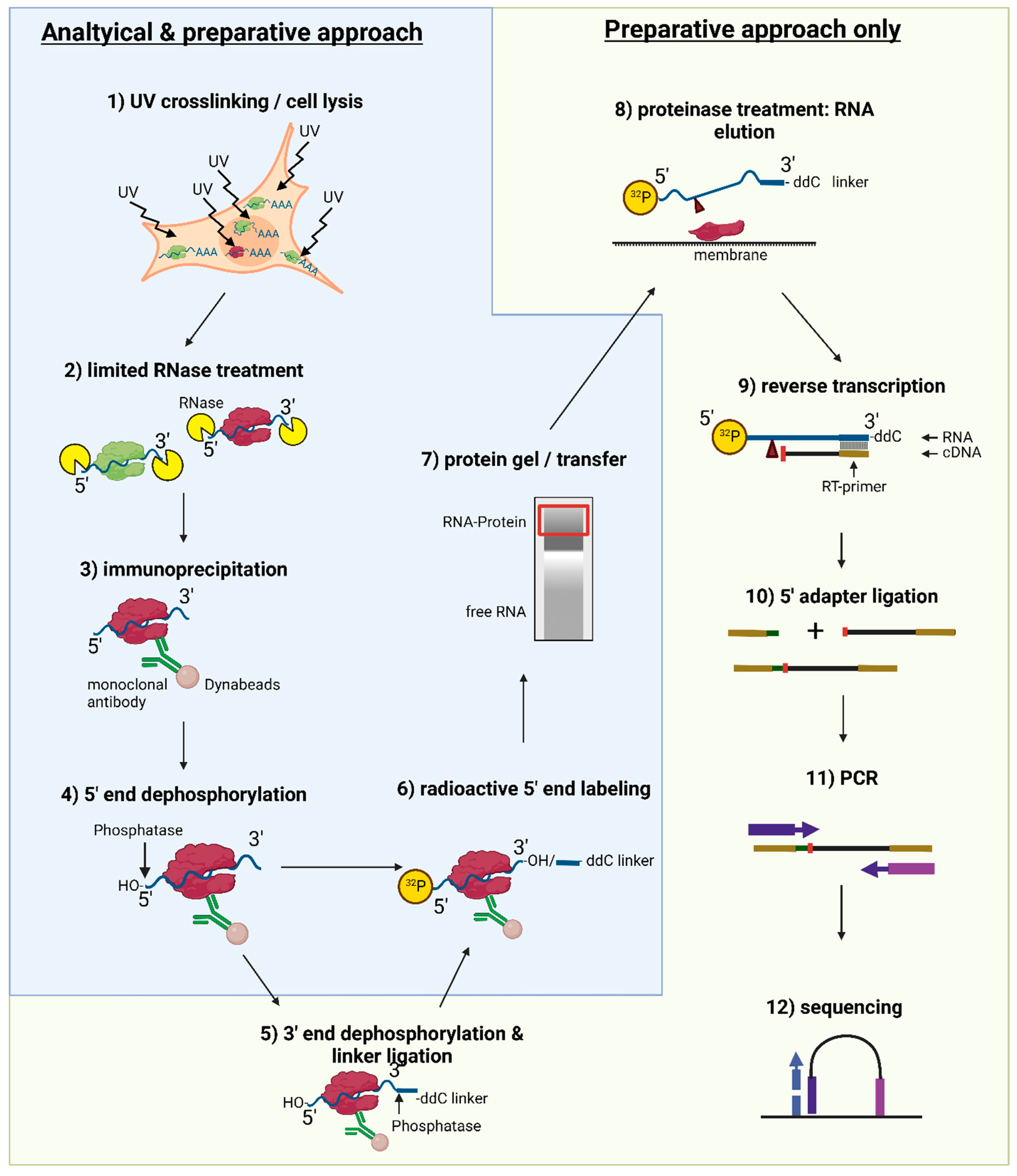

-1.jpg)








-2.jpg)